Gene expression Protein synthesis
Gene expression Protein synthesis
Dogma molecular biology
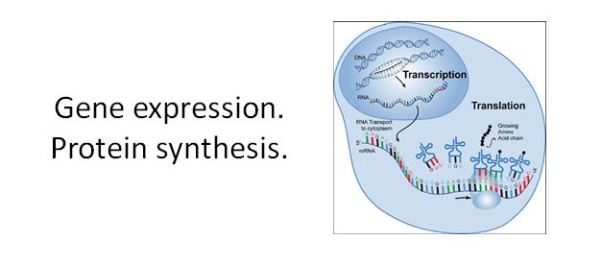
Gene expression is described by central dogma – two-step process denoted DNA → RNA → protein.
Gene expression is the process by which cells convert DNA sequence information to RNA and then decode the RNA information to the amino-acid sequence of a polypeptide.

Genes – the functional segments of DNA which code for the transfer of genetic information.
The genetic code by which DNA stores the genetic information consists of “codons” of three nucleotides.
With four possible bases, the three nucleotides can give 43 = 64 different possibilities.
The genetic code consists of 64 triplets of nucleotides. These triplets are called codons.
- But only 61 triplets or codons code for 20 amino acids used in the synthesis of proteins
- and 3 stop codons (aka nonsense codons or terminator codons) UAA, UAG, UGA.
Properties of genetic code:
1.The Genetic code is composed of nucleotide triplets.
2.The genetic code is degenerate
3.The genetic code is non-overlapping.
4.The genetic code is ordered (non-ambiguous).
5.The genetic code is comma-free.
6.The genetic code contains start and stop codons.
7.The genetic code is nearly universal
Properties of genetic code
1. The genetic code is triplet.
The Genetic code is composed of nucleotide triplets.
Three nucleotides in mRNA specify one amino acid in the polypeptide product;
thus, each codon contains three nucleotides.
2.The genetic code is degenerate, or redundant.
All but two (met, trp) of the amino acids are specified by more than one codon (from 2 to 6 ). For instance serine is encoded by six codons, glycine by four and lysine by two.
The first two letters seem to be the most important the third one tends to be interchangeable.
3.The genetic code is non-overlapping
Each nucleotide in mRNA belongs to just one codon
It means that the same letter does not take part in the formation of more than one codon.
4.The genetic code is ordered (non-ambiguous).
A particular codon will always code for the same amino acid.
It may also be that the same amino acid may be coded by two different codons.
However, when one codon codes for two amino acids, it is called ambiguous.
5.The code is comma less (comma-free):
The genetic code is without comma i.e. no punctuations are required between the two codons.
There are no demarcating signals between two codons.
This result is continuous coding of amino acid without interruption.
The reading frame is set at the beginning of the gene. Frame shift mutations can be caused by the ADDITION or DELETION of only one or two bases. Everything downstream is misread.
6.The genetic code contains «start» and «stop» codons.
Specific codons are used to initiate and to terminate polypeptide chains.
One codon, AUG serves two related functions:
- it signals the START of translation
- it codes for the incorporation of the amino acid methionine (Met) into the growing polypeptide chain
The codons UAA, UAG, and UGA – the STOP codons, which cause termination of translation by the ribosome.
7.The genetic code is nearly universal
With minor exceptions, the codons have the same meaning in all living organisms, from viruses to humans.
For instance, mitochondria have an alternative genetic code with small variations.
The reading of mRNA is always in the same direction 5’ to 3’ (the same way as transcription and replication).
The genetic code can be expressed as either RNA codons or DNA codons.
Different types of RNA are involved in the realization of genetic information and regulation of gene expression
Messenger RNA (mRNA)
- Сode for proteins
- Long Straight chain of Nucleotides
- Made up of 500 to 1000 nucleotides long
- Made in the Nucleus
- Copies DNA & leaves through nuclear pores
- Contains the Nitrogen Bases A, G, C, U ( no T )
- Carries the information for a specific protein
- Sequence of 3 bases called codon
- AUG – methionine or start codon
- UAA, UAG, or UGA – stop codons
Transfer RNA (tRNA)
- Clover-leaf shape
- Single stranded molecule with attachment site at one end for an amino acid
- Opposite end has three nucleotide bases called the anticodon
The tRNA carries a specific amino acid from the amino acid pool to the mRNA on the ribosomes to form a polypeptide, hence its name. The tRNAs form about 15% of the total RNA of a cell. Its molecule is the smallest and has the form of a cloverleaf. It has four regions:
- Acceptor stem – Carrier End: This is the 3 end of the molecule. Here a specific amino acid becomes attached. The tRNA molecule has a base triplet CCA with OH group at the tip. The COOH of amino acid joins the OH group.
- Anticodon – Recognition End: It is the opposite end of the molecule. It has 3 unpaired ribonucleotides. The bases of these ribonucleotides are complementary bases of the triplet found on mRNA chain called a codon. This triplet base sequence in tRNA is called as an anticodon. The anticodon binds with the codon at the time of translation.
- D loop – Enzyme Site: It is on one lateral side of the molecule. It is meant for a specific charging enzyme (aminoacyl tRNA synthetase) which catalyses the binding of a specific amino acid to tRNA molecule.
- TΨC loop – Ribosome Site: It is on the other lateral side of the molecule. It is meant for attachment to a ribosome.
Ribosomal RNA (rRNA) and ribosome
- rRNA is a single strand 100 to 3000 nucleotides long
- Globular in shape
- Made inside the nucleus of a cell
- Associates with proteins to form ribosomes
- Form the basic structure of the ribosome and catalyze protein synthesis
- Site of protein Synthesis
Differences Between RNA Types
| Features | Ribosomal RNA (rRNA) | Messenger RNA (mRNA) | Transfer RNA (tRNA) |
| 1. Percentage of cell’s total RNA | About 80 | About 5 | About 15 |
| 2. Length of molecule | Variable | Longest | Shortest |
| 3.Shape of molecule | Greatly coiled | Linear | Clover leaf- like, folded into L-shape |
| 4. Types | Six | Numerous | About 60 |
| 5. Role | Form greater part of ribosome’s | Carry information from DNA | Carry amino acids to mRNA codons |
| 6. Life | Long, used again and again in translation | Very short, 2 minutes to 4 hours, degraded after translation | Long, used again and again in translation |
Other types of RNAs
- snRNAs – small nuclear RNAs, function in a variety of nuclear processes, including the splicing of pre-mRNA
- snoRNAs – small nucleolar RNAs, used to process and chemically modify rRNAs
- miRNAs – microRNAs, regulate gene expression typically by blocking translation of selective mRNAs
- siRNAs – small interfering RNAs, turn off gene expression by directing degradation of selective mRNAs and the establishment of compact chromatin structures
- Other noncoding RNAs -function in diverse cell processes, including telomere synthesis, X-chromosome inactivation, and the transport of proteins into the ER
Stages of synthesis of protein:
1.Transcription
2.Translation
Transcription
- All types of RNA are synthesized from the DNA template on the principle of complementarity.
- This process is called transcription
- The enzymes that perform transcription are called RNA polymerases
Types of RNA polymerase
- There are three different types of RNA polymerase in eukaryotic cells (bacteria have only one):
- RNA pol I – transcribes the genes that encode most of the ribosomal RNAs (rRNAs)
- RNA pol II – transcribes the messenger RNAs
- RNA pol III – transcribes the genes transfer RNAs, one small rRNA, other small regulatory RNA molecules.
Sense and anti-sense strands of DNA
- The template for transcription is only one strand of DNA. It is called template or antisense.
- The second chain which is not involved in the transcription called coding or sense, has the same base sequence as mRNA(only with uracil instead of thymine)
Basic structure of a protein-coding gene
Transcription unit – promoter , RNA-coding sequence and terminator regions of a gene.
The promoter is upstream of the coding sequence, the terminator downstream . The coding sequence at nucleotide +1
Organization of eukaryotic gene.
- A promoter is a region of DNA that initiates transcription of a particular gene. Promoters are located near the transcription start sites of genes, on the same strand and upstream on the DNA (towards the 5’ region of the sense strand). Promoters can be about 100–1000 base pairs long.
- Core promoter – Includes the transcription start site (TSS) and elements directly upstream:
1) A binding site for RNA polymerase
2) General transcription factor binding sites, e.g. TATA boxes – – direct RNA polymerase to the correct initiation site for transcription.
- Proximal promoter – the proximal sequence upstream of the gene that tends to contain primary regulatory elements
RNA-coding sequence
Coding sequence of DNA present in mature messenger RNA;
DNA initially transcribed to messenger RNA consists of coding sequences (exons) and non-coding sequences (introns).
Introns are spliced out of the messenger RNA prior to translation, leaving only the exons to ultimately encode the amino acid product.
Splice site – specific base sequences at the junction between introns and exons. Cell can identify which bases should be saved and which should be removed
Organization of prokaryotic gene
Polycistronic model

- Prokaryotic cells have linear sequences of DNA called operons. An operon contains one or more structural genes which are generally transcribed into one polycistronic mRNA (a single mRNA molecule that codes for more than one protein).
- The operon is composed of a promoter sequence, followed by an operator gene, followed by one or more structural genes.
Organization of prokaryotic gene
An operon is made up of 3 basic DNA components:



1)Promotor – a nucleotide sequence that enables a gene to be transcribed. The promoter is recognized by RNA polymerase, which then initiates transcription.
2)Operator – a segment of DNA that a repressor binds to. In the case of a repressor, the repressor protein physically obstructs the RNA polymerase from transcribing the genes.
3)Structural genes– the genes that codes for proteins.
Regulator – These genes control the operator gene in cooperation with certain compounds called inducers and corepressors present in the cytoplasm. The regulator gene codes for and produces a protein substance called repressor. The repressor substance combines with the operator gene to repress its action. A regulator gene controls an operon, but is not the part of the operon.
Differences between the gene structure in prokaryotes and eukaryotes
1, A classical operon, found in prokaryotes (such as bacteria). Several genes, often functionally related, form a tight cluster on the genome. Operons are under the control of regulatory elements (promoter, operator) and factors that bind to these elements. Transcription of the cluster results in a single molecule, a multi-gene transcript of messenger RNA, which codes for several proteins and is directly translated into distinct protein products.
2, An operon in the nematode (eukaryotic organism). The initial multi-gene transcript is split up into separate messenger RNAs. The processing of their transcripts into proteins differs from that of prokaryotic operons.
Stages of transcription
1.Initiation
2.Elongation
3.Termination
Initiation
- The first step in transcription is initiation, when the RNA pol binds to the DNA upstream (5′) of the gene at a specialized sequence called a promoter.
- RNA polymerase unwinds and unzips DNA double strand, attaches to promoter region of gene, which marks the beginning point for transcription
Elongation:
RNA polymerase synthesizes a complementary RNA strand from nucleoside triphosphate taken from the surrounding solution in 5’ to 3’ direction
- It adds nucleoside triphosphates using base pairing rules:
A = U T = A G ≡ C C ≡ G
Termination
RNA polymerase reaches termination region of the gene, which marks the end of the coding sequence
- terminates transcription by releasing both DNA and RNA
Processing of pre-mRNA involves the following steps:
1.Capping – add 7-methylguanylate (m7G) to the 5′ end.
2.Polyadenylation – add a poly-A tail to the 3′ end.
3.Splicing – remove introns and join exons.
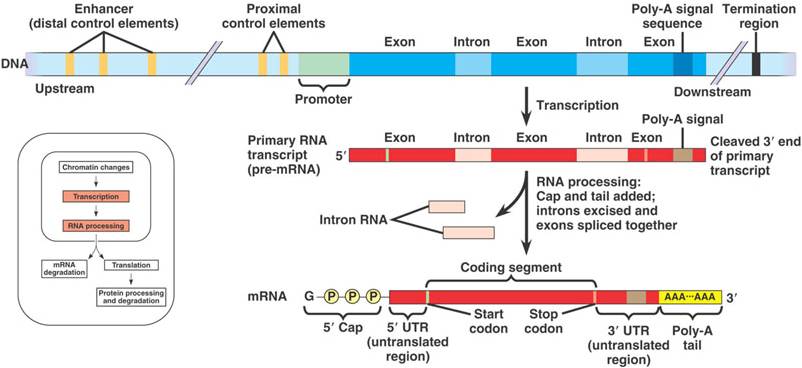

RNA Processing (maturation of pre-mRNA to mRNA): pre-mRNA → mRNA
- All the primary transcripts produced in the nucleus must undergo processing steps to produce functional RNA molecules
- Most human genes are divided into exons and introns.
- Primary transcripts (pre-mRNA) contains exons and introns.
- The exons are the sections that are found in the mature transcript (messenger RNA), while the introns are removed from the primary transcript by a process called splicing
Splicing
- the eukaryotic RNA needs the removal of introns to form mature mRNA and that this process is called splicing
- The removal of introns and splicing of exons is done by spliceosomes. These are a complexes of 5 snRNA molecules and some 145 different proteins.
- The introns in most pre-mRNAs begin with a GU and end with an AG (Splice site).
RNA processing steps
- Both ends of eucaryotic mRNAs are modified: by capping on the 5 end and by polyadenylation of the 3 end.
Alternative Splicing
- Alternative splicing of introns to form different mRNAs from the same gene.
- Alternative Splicing of exons allows one gene to make several different mRNAs, depending on which exons are included in the final message. Hence, one “gene” may code for a large number of different products. Some genes are known to make close to 100 different transcripts based on splicing patterns.
Transcription occurs when DNA acts as a template for mRNA synthesis. (49)
Translation occurs when the sequence of the mRNA codons determines the sequence of amino acids in a protein.
Translation
- Translation is the process of decoding the mRNA into a polypeptide chain
- In the cytoplasm, a ribosome attaches to the mRNA (three bases or 1 codon) and translates its message into a polypeptide
Ribosomes
- Made of a large (heavy) and small (light) subunit
- Composed of rRNA (40%) and proteins (60%)
- Have two sites for tRNA attachment – P and A
Stages of translation (protein Synthesis)
- Initiation
- Elongation
- Termination
- Protein modifications
Note: The same mRNA may be used hundreds of times during translation by many ribosomes before it is degraded (broken down) by the cell
Translation is initiated when an mRNA molecule, which has a ribosome-binding site in its 5′ UTR, binds the complementary sequence in the rRNA molecule that is part of the small ribosomal subunit, with the help of initiation factor proteins.
Another initiation factor assists in the binding of a charged tRNA molecule to the start codon.
This tRNA is formylmethionine in bacteria (the formyl group is later removed) and methionine in eukaryotes.
The large ribosomal subunit binds, with the initiatior tRNA in the P site.
Translation Elongation
Elongation begins when the next aminoacyl tRNA occupies the aminoacyl (A) site.
The amino acid shown here is glutamic acid, but any amino acid can be the second amino acid specified by a particular mRNA.
The peptidyltransferase center in the large subunit catalyzes the formation of a peptide bond bewteen the amino acid in the P site and the amino acid in the A site.
This reaction is catalyzed by ribosomal RNA (a ribozyme) rather than by a protein enzyme.
Once the peptide bond is formed, the mRNA is moved through the ribosome to place the tRNA with the growing peptide chain in the peptidyl (P) site.
A new aminoacyl tRNA is then free to occupy the aminoacyl site.
The next peptide bond is synthesized.
The mRNA is moved again, to place the tRNA with the growing peptide chain in the peptidyl (P) site.
Translation Termination
When the stop codon is reached, a release factor protein whose structure resembles that of a tRNA enters the A site.
Hydrolysis of the bond between the aminoacyl tRNA and the carboxy terminus of the last amino acid releases the peptide.
The tRNAs are also released
The ribosome separates from the mRNA and the two subunits of the ribosome dissociate.
Step 1- Initiation
- mRNA enters the cytoplasm
- mRNA transcript start codon AUG attaches to the small ribosomal subunit
- Small subunit attaches to large ribosomal subunit
Step 1- Initiation
- tRNAs, each carrying a specific amino acid, pair up with the mRNA codons inside the ribosomes. Base pairing (A-U, G-C) between mRNA codons and tRNA anticodons determines the order of amino acids in a protein.
- A complementary tRNA molecule with its attached amino acid (f-methionine) base pairs via its anticodon UAC with the AUG on the mRNA in the first position P.
Step 1- Initiation
- Another tRNA base pairs with the other three mRNA bases in the ribosome at position A.
- The enzyme peptidyl transferase forms a peptide bond between the two amino acids.
- The first tRNA (without its amino acid) leaves the ribosome.
Step 2- Elongation
- The ribosome moves along the mRNA to the next codon (three bases).
- The second tRNA molecule moves into position P.
- Another tRNA molecule pairs with the mRNA in position A bringing its amino acid.
- A growing polypeptide is formed in this way until a stop codon is reached.
Step 3– Termination
- A stop codon on the mRNA is reached and this signals the ribosome to leave the mRNA. The primary structure of protein (the sequence of its amino acids) is obtained.
Step 4- Modification
Most proteins undergo post-translational modifications.
Protein modification is the process by which some proteins from the rough ER are altered within the Golgi apparatus in order to be targeted to their final destinations.
The complete structure of functioning proteins involves more than polypeptide chains in the four levels of structure.
1.The primary structure is the sequence of amino acids.
2.Protein secondary structure refers to common repeating elements present in proteins. There are two basic components of secondary structure: the alpha helix and the beta-pleated sheet.
3.Tertiary structure is the full three-dimensional, folded structure of the polypeptide chain.
4.Quaternary structure is the spatial arrangement of two or more polypeptide chains. This structure may be a monomer, dimer, trimer, etc.
Step 4- Modification The stages of protein modifications:
- N- and C-terminal modifications
- Proteolytic Cleavage
- Folding of proteins (Secondary and Tertiary structure)
- Crosslinking (Quaternary structure)
N- and C-terminal modifications – addition of groups (methyl-, acetyl-, glyco-, phospho–):
- acetylation (N-terminal) – reaction that introduces an acetyl functional group into protein. Proteins are typically acetylated on lysine residues and this reaction relies on acetyl-coenzyme A as the acetyl group donor.
- methylation (N-terminal) – denotes the addition of a methyl group to protein. It takes place on arginine or lysine amino acid residues in the protein sequence.
- amidation (C-terminal) – the addition of an amide group from a glycine to a proteinamino acid (amidationof peptides (e.g., hormones) sometimes occurs at the C-terminus).
- phosphorylation – the addition of a phosphate group to proteins,
- glycosylation – attachment glycansto proteins.
- and other
Proteolytic Cleavage – deletion of parts to make a finished protein
- Frequently, the N-terminal methionine is not present in mature proteins
- Most proteins undergo proteolytic cleavage following translation. The simplest form of this is the removal of the initiation methionine. Many proteins are synthesized as inactive precursors that are activated under proper physiological conditions by limited proteolysis. Pancreatic enzymes and enzymes involved in clotting are examples of the latter. Inactive precursor proteins that are activated by removal of polypeptides are termed proproteins.
Protein folding
- Protein foldingis the process by which a protein structure assumes its functional shape or conformation.
- It is the physical process by which a polypeptide folds into its characteristic and functional three-dimensional structure from random coil.
Crosslinking
- Crosslinking is the process of chemically joining two or more molecules by a covalent bond.
- Attachment between two groups on a single protein results in intramolecular crosslinks that stabilize the protein tertiary or quaternary structure.
All these native modifications is extremely important because they may alter physical and chemical properties, folding, conformation distribution, stability, activity, and consequently, function of the proteins.
Control of an expression of genes – On and Off Genes
Clever mechanisms turn genes off and on so that they only function when there is a need for their services.
Genetic expression is the process where genotypes coded in the genes are exhibited by the phenotypes of the individuals.
Inducible and repressible system


The Discovery of the lac Operon
The discussion of prokaryotic gene expression focuses on the lac operon.
The lac operon of E. coli, originally investigated by Francois Jacob and Jacques Monod in the 1961, remains one of the best-understood models of gene regulation.
The model was based on their study of the genes in E. coli that code for enzymes that affect the breakdown of lactose.
Bacteria adapt to changes in their surroundings by using regulatory proteins to turn groups of genes on and off in response to various environmental signals.
The DNA of Escherichia coli is sufficient to encode about 4000 proteins, but only a fraction of these are made at any one time. E. coli regulates the expression of many of its genes according to the food sources that are available to it.
The lac operon in Escherichia coli.
Function – to produce enzymes which break down lactose (milk sugar)
ØLactose is not a common sugar, so there is not a great need for these enzymes
ØWhen lactose is present, they turn on and produce enzymes
Two components – repressor genes and functional genes
Three functional genes
The lac operon contains three structural genes, which encode enzymes that function in the metabolism of lactose.
- lacZ produces B-galactosidase. This enzyme hydrolyzes the bond between the two sugars, glucose and galactose
- lacY produces permease. This enzyme spans the cell membrane and brings lactose into the cell from the outside environment. The membrane is otherwise essentially impermeable to lactose.
- lacA produces B-galactosidase transacetylase. The function of this enzyme is still not known.
Promoter (P) – aids in RNA polymerase binding
Operator (O) – “on/off” switch – binding site for the repressor protein
The operator is a short region of DNA that lies partially within the promoter and that interacts with a regulatory protein that controls the transcription of the operon.
Repressor (lacI) gene
Repressor gene (lacI) – produces repressor protein (two binding sites, one for the operator and one for lactose). Regulatory genes are not necessarily close to the operons they affect.
The repressor protein is under allosteric control – when not bound to lactose, the repressor protein can bind to the operator
When lactose is present, an isomer of lactose, allolactose, will also be present in small amounts. Allolactose binds to the allosteric site and changes the conformation of the repressor protein so that it is no longer capable of binding to the operator.
The lac operon Escherichia coli.
The Lac regulatory protein is called a repressor because it keeps RNA polymerase from transcribing the structural genes.
Thus the Lac repressor inhibits transcription of the lac operon.
Operation – If lactose is not present:
the repressor gene produces repressor, which binds to the operator.
This blocks the action of RNA polymerase, thereby preventing transcription.
Operation – if lactose is present:
- the repressor gene produces repressor, which has a site for binding with allolactose.
- The allolactose /repressor compound is incapable of binding/ the operator, so the RNA polymerase is uninhibited
- once the concentration of lactose decreases, the repressor-allolactose complex falls apart and transcription is again inhibited.
- The lac operon is an example of an inducible operon – it is normally off, but when a molecule called an inducer is present, the operon turns on.
- The trp operon is an example of a repressible operon – it is normally on but when a molecule called a repressor is present the operon turns off.
Gene Regulation in Eukaryotes
- The latest estimates are that a human cell, a eukaryotic cell, contains approximately 35,000 genes.
- Some of these are expressed in all cells all the time. These so-called housekeeping genes are responsible for the routine metabolic functions (e.g. respiration) common to all cells.
- Some are expressed as a cell enters a particular pathway of differentiation.
- Some are expressed all the time in only those cells that have differentiated in a particular way. For example, a plasma cell expresses continuously the gene for the antibody it synthesizes.
- Some are expressed only as conditions around and in the cell change. For example, the arrival of a hormone may turn on (or off) certain genes in that cell.
There are several methods used by eukaryotes.
Transcription Control
The most common type of genetic regulation
Turning on and off of mRNA formation
Post-Transcriptional Control
Regulation of the processing of a pre-mRNA into a mature mRNA
Translational Control
Regulation of the rate of Initiation
Post-Tranlational Control
Regulation of the modification of an immature or inactive protein to form an active protein
Regulatory sequences:
- Enhancers,
- Silencers,
- Insulators
Enhancers
Some transcription factors (“Enhancer-binding protein”) bind to regions of DNA that are thousands of base pairs away from the gene they control. Binding increases the rate of transcription of the gene.
Silencers
Regulatory sequences with similar characteristics, but the opposite effect, exist. These are called silencers.
Silencers are control regions of DNA that, like enhancers, may be located thousands of base pairs away from the gene they control. However, when transcription factors bind to them, expression of the gene they control is repressed.
Insulators
As you can see above, enhancers can turn on promoters of genes located thousands of base pairs away.
What is to prevent an enhancer from inappropriately binding to and activating the promoter of some other gene in the same region of the chromosome?
One answer: an insulator.
Insulators are stretches of DNA (as few as 42 base pairs may do the trick) located between the
- Enhancer(s) and promoter or
- Silencer(s) and promoter
of adjacent genes or clusters of adjacent genes.
Their function is to prevent a gene from being influenced by the activation (or repression) of its neighbors.
Major rearrangements of at least one set of genes occur during immune system differentiation.
B lymphocytes produce immunoglobins, or antibodies, that specifically recognize and combat viruses, bacteria, and other invaders.
Each differentiated cell produces one specific type of antibody that attacks a specific invader.
Functional antibody genes are pieced together from physically separated DNA regions.
Each immunoglobin consists of four polypeptide chains, each with a constant region and a variable region, giving each antibody its unique function.
As a B lymphocyte differentiates, one of several hundred possible variable segments is connected to the constant section by deleting the intervening DNA.
The random combinations of different variable and constant regions create an enormous variety of different polypeptides, which combine with others to form complete antibody molecules.
As a result, the mature immune system can make millions of different kinds of antibodies from millions of subpopulations of B lymphocytes.
Gene expression Protein synthesis
Gene expression Protein synthesis